David Josephy, PhD, Executive Director of GIST Sarcoma LRG Canada
 I attended the CTOS (Connective Tissue Oncology Society) Annual Meeting in Vancouver, British Columbia, Nov. 16-19, representing GIST Sarcoma Life Raft Group Canada. CTOS is an international society of physicians and scientists with a primary interest in sarcomas (tumors of connective tissues). The goal of the society is to advance the care of patients with connective tissue tumors and to increase knowledge of all aspects of the biology of these tumors, including basic and clinical research. Joining Life Raft Group Executive Director, Sara Rothschild and Senior Director of Data Management and Research, Denisse Montoya, we were able to hear information about the latest research in GIST and other cancers, and to network with colleagues from around the globe to explore potential collaborations.
I attended the CTOS (Connective Tissue Oncology Society) Annual Meeting in Vancouver, British Columbia, Nov. 16-19, representing GIST Sarcoma Life Raft Group Canada. CTOS is an international society of physicians and scientists with a primary interest in sarcomas (tumors of connective tissues). The goal of the society is to advance the care of patients with connective tissue tumors and to increase knowledge of all aspects of the biology of these tumors, including basic and clinical research. Joining Life Raft Group Executive Director, Sara Rothschild and Senior Director of Data Management and Research, Denisse Montoya, we were able to hear information about the latest research in GIST and other cancers, and to network with colleagues from around the globe to explore potential collaborations.
The meeting included a large number of presentations and posters on GIST research, which is encouraging. The Life Raft Group presented an original research poster entitled Phase I Results from a Multi-Phase Comprehensive Genomic Sequencing Tumor Study in Gastrointestinal Stromal Tumor and was also part of research presented on GIST & Other Sarcomas with Actionable Targets with Andrea Napolitano, MD, PhD et al. Several oral presentations highlighted promising findings.
Presentation #1:

Dr. Xiaolan Feng
The next study, entitled Refining Prognosis in Localized GIST: Clinical Significance of PTEN Low Expression and Gene Loss was by led by Dr. Xiaolan Feng. Dr. Feng is a medical oncologist. She was practicing in Victoria, B.C., until recently, and has now moved back to Calgary, Alberta (Tom Baker Cancer Centre). She has received some funding from LRG Canada.
Objective: To investigate the use of PTEN biomarker to improve prognostic stratification in patients with localized GIST.
PTEN is a tumor suppressor gene and regulates the PI3K/Akt/mTOR signaling pathway. (See: Stefano and Giovanni, The PTEN tumor suppressor gene in soft tissue sarcoma, Cancers (Basel) 2019 doi: 10.3390/cancers11081169) PTEN loss and activation of the PI3K/Akt/mTOR pathway is frequently seen in advanced/ imatinib-resistant GIST.
Methods: They measured PTEN expression (RNA levels) and PTEN gene copy number (normal vs heterozygous or homozygous loss) on two independent cohorts: GIST-60 (n=60) and GIST-100 (n=100); frozen tumor samples were obtained from primary tumor resection.
Results: PTEN expression (RNA level) was classified as either low or high, relative to the mean.
As stated in the published paper, “The transcriptional profiling in the GIST-60 cohort revealed a significantly lower PTEN expression in patients with local and metastatic relapse compared with those with no documented recurrence”. PTEN low expression was also significantly associated with poor disease-free survival (DFS). DNA analysis showed heterozygous loss of PTEN in approximately 10% of the patients in each cohort and this parameter was also associated with poor DFS compared vs patients with PTEN normal status.
Conclusion: PTEN low expression/gene loss is an independent significant prognostic factor, and a promising component to strengthen the clinical prognostic tools in patients with localized GIST.
In the publication, they suggest that “use of a further validated PTEN immunohistochemistry assay could allow stratification of clinically high-risk patients to participate in clinical trials to investigate the additional value of mTOR inhibitors to standard adjuvant imatinib, and clinically low-/intermediate-risk patients to undergo intense surveillance.” On her final slide, Dr Feng stated that further validation (e.g. by IHC) is warranted and that “collaboration with GIST Life Raft and other stakeholders is underway.”
This is an exciting opportunity!
Presentation #2:
Steven Bialick, Jonathan Trent, et al. (University of Miami/Sylvester Comprehensive Cancer Center) presented Increased Progression Free Survival (PFS) in Advanced GIST patients Treated with Sunitinib After Circulating Tumor DNA Detection of a KIT Exon 13 Resistance Mutation.

Dr. Steve Bialick
In this study, they performed a retrospective analysis of ctDNA Next-Generation Sequencing (“Guardant360″) results from GIST patients (n=104) who had progressed on first line imatinib. Of this group, 64 patients had primary KIT alterations; 40 did not and were not analyzed further. Among the 64, 46 patients had known imatinib-resistance mutation in either KIT exon 13 or exon 17 or both. Among the exon 13 imatinib-resistance mutations (28 patients), 25 patients showed only the common V654A mutation. Among the exon 17 imatinib-resistance mutations (35 patients), 20 patients showed only an exon 17 mutation; 15 showed both exon 13 and 17 imatinib-resistance mutations. 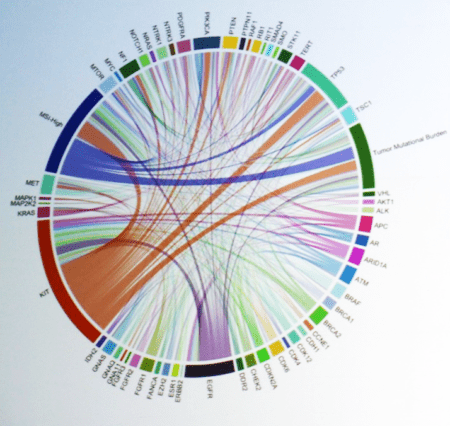
Bialick showed a colorful circular graphical representation of 57 different mutations (or biomarkers) that were detected in the 104 patient samples, with “chords” representing the co-occurrence of multiple genes/ biomarkers in the same patient.
Among these genes were KIT and also: PDGFRA; PTEN; TP53; RB1; VHL; ATM; BRCA1; BRCA2; CDK4; EGFR; FANCA; KRAS; NRAS; MET; MTOR; MYC; NTRK1; NTRK3; and other cancer-related genes. He showed Kaplan-Meier PFS plots for (a) the 25 patients with only the exon 13 V654A imatinib-resistance mutation: and (b) the 35 patients with exon 17 imatinib-resistance mutations. For the exon 13 V654A group, 25 patients, median PFS was much longer for patients treated with sunitinib (11.3 months) than for patients treated with regorafenib (6.2 months) or other drugs (3.2 months). For the exon 17 group, 35 patients, there were no significant differences among the three treatment sub-groups, with median PFS about 6 months for sunitinib, regorafenib, or other drugs.
This suggests that ctDNA detection of secondary mutations could, as we would hope, be used as guidance in choosing second- and further-line therapies. He concluded that “ctDNA-guided therapy warrants evaluation in a prospective clinical trial”.
Presentation #3:
A novel prognostication system for patients with high-risk KIT exon 9-mutated GIST receiving adjuvant imatinib, Andrea Napolitano, Sebastian Bauer, Heikki Joensuu, Denisse Montoya, et al.

Dr. Andrea Napolitano
Objective: Recurrence risk evaluation in patients with resected GISTs harbouring KIT exon 9 mutations is commonly performed using systems not incorporating the mutational status, such as the Armed Forces Institute of Pathology (AFIP) criteria or prognostic “heat-maps”. GIST patients with a high-risk of recurrence receive adjuvant treatment with imatinib. No prognostication systems specific to high-risk KIT ex9 GIST patients receiving adjuvant imatinib exist.
They “retrospectively collected baseline clinical and pathological characteristics and relapse-free survival (RFS) data of 231 high-risk exon-9 GIST patients who received adjuvant treatment with imatinib.” A prognostic nomogram was developed. They found significant negative prognostic roles for: male; older at diagnosis; largest maximum tumour dimension; extra-gastric site; mitotic count. The allocated dose of adjuvant imatinib (400 vs 800 mg/day) was not a prognostic factor. They developed a prognostic nomogram including all of the clinical variables specific for this population. This prognostic nomogram should be applicable right away, to evaluate risk of recurrence and guide the decision on whether to take adjuvant imatinib, for exon 9 patients.
Presentation #4:

Dr. Joanna Przybyl
The next presentation was from Dr. Joanna Przybyl and includes Dr. Matt van de Rijn as a contributing researcher.
Multi-omic Integrative Profiling of miniGISTs and Clinically Relevant GISTs Throughout Progression – Dr. Przybyl is a PhD research scientist. She trained with Dr. Piotor Rutkowski in Poland and with Dr. Van De Rijn at Stanford. Last year, she started as Assistant Professor at McGill University in Montreal. She has received support from LRGC and she spoke at our recent LRG Canada AGM in Toronto. This presentation was based on her research at Stanford.
Quoting from the Abstract:
Objective: GISTs <1cm (miniGISTs) occur frequently, with the highest reported incidence of approximately 30% in the general population. In contrast, clinically relevant malignant GISTs are very rare. It remains unknown which molecular programs are involved in the early stages of malignant transformation in GISTs. Given the great variation in incidence, it is not even clear whether miniGISTs can be considered clinical precursors for malignant GISTs We performed a multiplatform in-depth molecular analysis of miniGISTs and overt GISTs at different stages of progression to identify molecular programs that contribute to development of clinically relevant GISTs.
Methods: We profiled DNA copy number alterations, DNA methylation and RNA expression of 59 primary untreated GISTs. These included 16 miniGISTs ( < 1cm), 20 small GISTs (1-5 cm) and 23 large GISTs (> 5 cm) We also performed targeted DNA sequencing of 400 cancer-related genes for 16/59 cases. We validated selected findings by immunohistochemistry and in situ hybridization.
Results: All miniGISTs included in our dataset carried mutations of KIT or PDGFRA genes, and 5 of 14 miniGISTs with DNA copy number data available showed loss of chromosomes 14, 1p, 15 or 13 that have been reported in clinically relevant GIST.
Our multi-omic analysis shows that the largest difference in molecular profiles between miniGISTs and overt GISTs is manifested on transcriptomic level, as opposed to DNA copy number or DNA methylation level. In comparison to miniGISTs, overt GISTs lose the expression of genes involved in myogenesis, matrisome and inositol phosphate pathway metabolism – a pathway that is known to be active in the interstitial cells of Cajal.
Transcriptomic (RNA) data shows that small GISTs first gain the enrichment of target genes regulated by ZFHX3 and ETS2 transcription factors and genes related to cell growth and motility In addition, overt GISTs show enrichment of genes involved in G2M checkpoint, mTORC1 signaling, mitotic cell spindle organization and genes regulated by E2F transcription factor that gradually increases with the size of the tumor. Through an integrative analysis of genomic, epigenomic and transcriptomic data, we identify 21 novel candidate tumor suppressor genes affected by frequent copy number loss and epigenetic silencing on chromosomes 14, 1p, 22, 15, 13 and 11.
Conclusion: Our results show that miniGISTs may present similar genomic alterations as small and large GISTs. Thus, development and progression of clinically relevant disease may be rather associated with the activation of specific transcriptomic programs that promote oncogenesis. Our study offers the first multi-omic characterization of GISTs, including DNA methylation profiles, that identifies novel tumor suppressor genes that may contribute to progression of GISTs. This study also presents the first in-depth analysis of multi-omic profiles of miniGISTs.”
Presentation #5
The final presentation in the GIST session was led by Patrick Schöffski et al., Leuven Cancer Institute, Leuven, Belgium titled, Anti-tumor effects of the novel kit mutant inhibitor M4205/IDRX-42 in GIST xenograft models.
First, a little background about IDRX-42:
From: Paul Lim, Blueprint Medicines; Tues. Aug. 2, 2022
I wanted to share with you the news that Blueprint Medicines has licensed a development candidate-stage KIT exon 13 inhibitor to IDRx, a clinical-stage biopharmaceutical company focused on purpose-built precision combination therapies in cancer.
First-in-human study of IDRX-42; Dana-Farber Cancer Institute, Boston;
Contact: Suzanne George, MD
Cancer Research 82, Supplement, 15 June 2022; Poster Presentations – Late-Breaking Abstracts
Abstract LB565: Efficacy of a highly potent and selective KIT V654A inhibitor for treatment of imatinib resistant GIST; Alexandra R. Grassian et al.
IDRX-42 is a novel ATP-competitive KIT inhibitor. Biochemical and in vitro assays show that KIT mutations in exons 11, 13, and 17 are sensitive to the inhibitor and that it has excellent kinase selectivity. The drug candidate has good human pharmacokinetics and no predicted brain penetration. A Phase 1 first-in-human trial has been clinical started.

Dr. Patrick Schöffski
From the Schöffski Abstract:
We tested the efficacy of M4205/IDRX-42, a novel specific KIT inhibitor with high activity towards the most relevant KIT mutations, in GIST xenograft models.
[Nude mice: The nude mouse is a naturally-occurring mutant that lacks a thymus and also lacks hair. The lack of a thymus causes T-cell deficiency, so the nude mouse is immunodeficient and can accept foreign tissues such as human tumors.]
NMRI nu/nu mice were transplanted with patient-derived GIST xenograft models UZLX-GIST9, resistant to both imatinib and sunitinib; with the dose-dependent imatinib- and sunitinib-sensitive models UZLX-GIST2B and UZLX-GIST25; and with the cell-line derived model GIST882. Mice were treated daily with vehicle (control), imatinib (100 mg/kg), avapritinib (5 mg/kg), sunitinib (20 mg/kg) or M4205/IDRX-42 (10 or 25 mg/kg).
Results: M4205/IDRX-42 (25 mg/kg) caused tumor volume shrinkage in all of the models. Antitumor activity of M4205 was superior to imatinib in UZLX-GIST9, -GIST2B and GIST882, and superior to sunitinib in -GIST25.
Conclusion: M4205/IDRX-42 has significant antitumor activity in patient- and cell line-derived GIST xenograft models.
Posters
There were over twenty posters relevant to GIST research. Highlighted here are several with the potential for rapid clinical impact. For further information about these posters, you may reach out to the authors or to patientregistry@liferaftgroup.org
Poster #96 2206814
AN OPEN-LABEL, PHASE 2 EFFICACY STUDY OF TEMOZOLOMIDE IN ADVANCED SUCCINATE DEHYDROGENASE-MUTANT/DEFICIENT GASTROINTESTINAL STROMAL TUMOR, Adam Burgoyne, MD, PhD1; Michael C. Heinrich, MD2; Margaret von Mehren, MD3; Jonathan C. Trent, MD, PhD4; Emily Messer, PhD1; Karen Messer, PhD1; Christian Metallo, PhD5; Jason K. Sicklick, MD, FACS6
(1) UC San Diego Moores Cancer Center, La Jolla, CA, (2) Oregon Health and Science University, Portland, OR, (3) Fox Chase Cancer Center, Philadelphia, PA, (4) University of Miami/Sylvester Comprehensive Cancer Center, Miami, FL, (5) Salk Institute, La Jolla, CA, (6) University of California, San Diego, San Diego, CA
Poster #100 2206886
COPY NUMBER ALTERATIONS IN CDKN2A AND CDKN2B ARE ASSOCIATED WITH WORSENED OVERALL SURVIVAL IN PATIENTS WITH GIST, Ashwyn K. Sharma, MD; Winta T. Mehtsun, MD, MPH; Jill P. Mesirov, PhD; Jason K. Sicklick, MD, FACS, University of California, San Diego, San Diego, CA
Poster #106 2206933
LAPAROSCOPY IN GASTRIC GIST: RETROSPECTIVE PRE-POST COMPARISON ANALYSIS OF SHORT TERM OUTCOMES ON A CONSECUTIVE SERIES OF 95 CASES FROM A TERTIARY REFERRAL CENTRE, Laura Ruspi, MD1; Laura Samà, Surgeon2; Manuela Cammelli, MD3; Ilaria Santori, MD4; Marta Tassan-Mangina, MD5; Federico Sicoli, MD6; Sonia Kumar, Surgeon5; Alice Laffi, MD5; Alexia Bertuzzi, MD5; Piergiuseppe Colombo, MD7; Lorenzo Salvatore Renne, MD8; Ferdinando Carlo Maria Cananzi, MD6; Vittorio Quagliuolo, MD9
(1) Sarcoma, Melanoma and Rare Tumors Surgery Unit, Humanitas Clinical and Research Center, IRCCS, Rozzano, 20089, Italy, Milano, Lombardia, Italy, (2) Humanitas Research Hospital, Milano, Lombardia, Italy, (3) Humanitas University, Rozzano, Lombardia, Italy, (4) Humanitas University, Rozzano, Emilia-Romagna, Italy, (5) Humanitas Research Hospital, Milan, Lombardia, Italy, (6) Sarcoma Surgery Unit, Humanitas Clinical and Research Center, IRCCS, Rozzano, 20089, Italy, Rozzano, Lombardia, Italy, (7) Pathology Unit, Humanitas Clinical and Research Center, IRCCS, Rozzano, 20089, Italy, Rozzano, Lombardia, Italy, (8) Humanitas Research Hospital, Rozzano, Lombardia, Italy, (9) Istituto Clinico Humanitas, Rozzano, Lombardia, Italy
Poster #108 2206788
NILOTINIB REVISITED: SALVAGE USE IN PATIENTS WITH SEVERE IMATINIB-TOXICITY, Johanna Falkenhorst, MD1; Thomas Mühlenberg, PhD1; Benjamin Fletcher1; Jürgen Treckmann, MD2; Moritz Kaths, MD2; Farhad Farzaliyev, MD3; Johannes Grüneisen, MD4; Rainer Hamacher, MD5; Tom Schulz6; Alina Teuber, MSc6; Hans-Ulrich Schildhaus, MD7; Daniel Rauh, MD6; Sebastian Bauer, MD8
(1) Department of Medical Oncology, Sarcoma Center, West German Cancer Center, University Duisburg-Essen, Medical School, Essen, Germany, Essen, Nordrhein-Westfalen, Germany, (2) Department of Visceral Surgery, Sarcoma Center, West German Cancer Center, University Duisburg-Essen, Medical School, Essen, Germany, Essen, Nordrhein-Westfalen, Germany, (3) Department of Plastic Surgery, BG Klinik Tübingen, Tübingen, Germany, Tübingen, Baden-Wurttemberg, Germany, (4) Institute of Radiology, University Duisburg-Essen, Medical School, Essen, Germany, Essen, Nordrhein- Westfalen, Germany, (5) Department of Medical Oncology, West German Cancer Center, University Hospital Essen, Essen, Nordrhein-Westfalen, Germany, (6) Faculty of Chemistry and Chemical Biology, TU Dortmund University, Dortmund, Germany, Dortmund, Nordrhein-Westfalen, Germany, (7) Institute of Pathology, West German Cancer Center, University Hospital Essen, Essen, Nordrhein-Westfalen, Germany, (8) Department of Medical Oncology, Sarcoma Center, West German Cancer Center, University Hospital Essen, University of Duisburg-Essen and German Cancer Consortium (DKTK), Partner Site University Hospital Essen, Essen, Germany, Essen, Nordrhein-Westfalen, Germany
Poster #121 2206900
USING IN VITRO MODELS TO PREDICT IMATINIB RESPONSES IN PDGFRA-MUTANT GASTROINTESTINAL STROMAL TUMOR, Homma M. Khosroyani, BS1; Ajia Town, BS1; Lillian R. Klug, PhD1; Johanna Falkenhorst, MD2; Sebastian Bauer, MD3; Michael C. Heinrich, MD4
(1) OHSU Knight Cancer Institute, Portland, OR, (2) Department of Medical Oncology, Sarcoma Center, West German Cancer Center, University Duisburg-Essen, Medical School, Essen, Germany, Essen, Nordrhein-Westfalen, Germany, (3) Department of Medical Oncology, Sarcoma Center, West German Cancer Center, University Hospital Essen, University of Duisburg-Essen and German Cancer Consortium (DKTK), Partner Site University Hospital Essen, Essen, Germany, Essen, Nordrhein-Westfalen, Germany, (4) Oregon Health and Science University, Portland, OR
Clinical trials:
Poster #316 2206547
FIRST-IN-HUMAN PHASE 1/1B STUDY OF IDRX-42, A NOVEL ORAL TYROSINE KINASE KIT INHIBITOR,
IN PARTICIPANTS WITH METASTATIC AND/OR UNRESECTABLE GASTROINTESTINAL STROMAL TUMORS, Suzanne George, MD1; George Demetri, MD2; Nick Lydon, PhD2; Vivek Kadambi, PhD2; Jessica Christo, MA2; Debbie Johnson, BS2; Patrick Schöffski, MD, MPH3
(1) Dana-Farber Cancer Institute, Boston, MA, (2) IDRx, Inc, Plymouth, MA, (3) Laboratory of Experimental Oncology, Department of Oncology, KU Leuven, Leuven Cancer Institute, Leuven, Vlaams-Brabant, Belgium
Poster #320 2206503
PEAK STUDY: A PHASE 3 RANDOMIZED, OPEN-LABEL, MULTICENTER CLINICAL STUDY OF BEZUCLASTINIB (CGT9486) AND SUNITINIB VERSUS SUNITINIB IN PATIENTS WITH GASTROINTESTINAL STROMAL TUMORS, Andrew J. Wagner, MD, PhD1; Sebastian Bauer, MD2; Michael C. Heinrich, MD3; Robin Lewis L. Jones, MD4; César Serrano, MD, PhD5; Margaret von Mehren, MD6; Neeta Somaiah, MD7; Tom Andor8; Benjamin Exter, PharmD8; Dirk Huebner, MD8; Zamaneh Mikhak, MD8; Shaunica Mitchell8; Kevin Moynihan8; Jessica Sachs, MD8; Lei Sun, PhD8; Jonathan C. Trent, MD, PhD9; William D. Tap, MD10
(1) Dana-Farber Cancer Institute and Harvard Medical School, Boston, MA, USA, Boston, MA, (2) Department of Medical Oncology, Sarcoma Center, West German Cancer Center, University Hospital Essen, University of Duisburg-Essen and German Cancer Consortium (DKTK), Partner Site University Hospital Essen, Essen, Germany, Essen, Nordrhein-Westfalen, Germany, (3) Oregon Health and Science University, Portland, OR, (4) Royal Marsden Hospital and Institute of Cancer Research, London, England, United Kingdom, (5) Vall D’hebron Institute of Oncology, Barcelona, Catalonia, Spain, (6) Fox Chase Cancer Center, Philadelphia, PA, (7) The University of Texas MD Anderson Cancer Center, Houston, TX, (8) Cogent Biosciences, Cambridge, MA, (9) University of Miami/Sylvester Comprehensive Cancer Center, Miami, FL, (10) Memorial Sloan Kettering Cancer Center and Weill Cornell Medical College, New York, NY, USA, New York, NY
The Life Raft Group Presented Original Research
Poster #110: Phase I Results from A Multi-Phase Comprehensive Genomic Sequencing Tumor Study In Gastrointestinal Stromal Tumor Patients.
PHASE I RESULTS FROM A MULTI-PHASE COMPREHENSIVE GENOMIC SEQUENCING TUMOR STUDY IN GASTROINTESTINAL STROMAL TUMOR PATIENTS, Denisse Montoya, MSc; Sahibjeet Kaur, MPH; Antoinette Pauwels; Olivia Regan; Katie Regan; Jim Hughes; Sara Rothschild, MPH, The Life Raft Group, Wayne, NJ

To see larger version of poster, click on image.
Objective: Gastrointestinal Stromal Tumors (GISTs) are the most common mesenchymal neoplasms of the gastrointestinal tract Most GISTs harbor a mutation located in either KIT or platelet-derived growth factor receptor alpha (PDGFRA) Over the last few decades, advances in GIST have led to the approval of multiple targeted therapies Imatinib, one of the frequently used therapies for GIST, has demonstrated low sensitivity among KIT/PDGFRA wildtype GISTs. Comprehensive genomic profiling (CGP) is used to optimize the therapy selection due to its ability to detect underreported pathogenic biomarkers and allowing a more tailored approach to therapy in both primary and metastatic settings Several targetable drivers including neurotrophic tyrosine receptor kinase (NTRK) fusions have been identified by using CGP and could benefit patients with FDA approved therapies Despite the advances of therapies and the known importance of CGP, the rate of CGP is relatively low in patients with GIST The present study reports results from Phase I of our multi-phase study, which will demonstrate the various genomic drivers that exist in this population and the potential impact of treatment trajectory for patients.
Methods: Next generation sequencing (NGS) based biomarker analysis was performed on tumor DNA and RNA focusing on wildtype GIST subgroups and included NTRK gene fusion Patient recruitment involved The Life Raft Group (LRG) GIST Patient Registry members, direct patient outreach via phone and email, social media, and physician referral with a proposed sample size of 255 patients to be tested at the end of the multiphase study Eligible patients were required to reside in the USA, have never received any form of genomic testing for their GIST diagnosis, be an active member of the LRG GIST Patient Registry, and have viable formalin-fixed, paraffin-embedded (FFPE) tissue slides or blocks available for testing. Patients who showed a wildtype KIT/PDGFRA result from a previous basic KIT/PDGFRA mutational test were also eligible The test featured a 20 gene panel including sequencing coding DNA of all the exons within the genes plus an additional 50 nucleotides at the 5’ and 3’ ends of each coding exon to detect splicing abnormalities Reports were shared with the patient and treating physician to aid in treatment selection Results from Phase I were analyzed to understand the impact within the patient cohort and potential therapy changes.
Results: 104 patients expressed interest in the study. 55 (53%) patients were qualified for testing while 49 (47%) patients were not qualified. Out of the 49 unqualified, 18 (37%) did not meet residency requirements, 19 (39%) did not have any tissue available for testing and 12 (24%) patients already had CGP performed but were unaware or had not reported results to the patient registry Tumor tissue from 55 eligible patients was processed through NGS Patient characteristics from this cohort are portrayed in the table below (Table 1) Out of the samples tested, 23 (42%) patients had a previous wildtype KIT/PDGFRA reported mutation and 32 (58%) patients never had received CGP 29 (53%) patients received a Test Not Per- formed (TNP), Quantity Not Sufficient (QNS) or showed results with Variants of Unknown Significance (VUS). Biomarkers were identified in 26 (47%) patients. Remarkably, NTRK ETV6-fusion positive was present in 2 (2/26; 8%). Half of the mutations detected were KIT mutations (13/26; 50%) PDGFRA mutation was present in 2 (8%) patients, VUS in EGFRA mutation was detected in one patient 3 (11%) patients showed an NF1 mutation, with a dual mutation of SDHB and NF1 detected in one patient SDHA mutation was present in 6 (23%) patients With these results, 11 (42%) patients had to change or adjust their medication to ensure the most effective treatment plan
Conclusion: Utilizing NGS based CGP is a crucial step in understanding and identifying relevant biomarkers including tumor growth drivers to aid in the treatment decision for optimal patient outcomes CGP provides relevant data for conducting a more accurate, individualized treatment plan for patients, and prevents patients from undergoing unnecessary therapies that will not work for their respective mutation Furthermore, this approach accelerates adoption of precision oncology and aids future drug discovery and development programs specifically for GIST patients. Genomic characterization is a critical point in the treatment course of countless GIST patients and should be considered the standard practice for GIST patients that qualify for oral chemotherapy treatment. Based on Phase I findings, a Phase II study is underway to explore additional genomic drivers that may exist in this population, and how targeted therapies can change the treatment trajectory and outcome for patients Phase II will expand CGP to feature a 648 gene DNA panel sequence, Microsatellite Instability (MSI) status, Tumor Mutational Burden (TMB), and full transcriptome analysis by RNA sequencing. Patient-matched germline DNA will also be explored to further understand the genomic landscape in GIST.
The conference resulted in a number of exciting possibilities for collaborative research. We look forward to new avenues of study from sarcoma researchers.
 Our Good News Holiday Campaign is the culmination of our year of “Time to Tell the Stories 2022.” In this series, we celebrate the connections, the celebrations, the events and milestones that we’ve been privileged to be a part of this year. Our 20th anniversary year is winding down and we are wrapping it up by spreading good news & gratitude throughout the season. Your financial donations and selfless volunteerism are what enable us to continue providing vital services to our GIST community.
Our Good News Holiday Campaign is the culmination of our year of “Time to Tell the Stories 2022.” In this series, we celebrate the connections, the celebrations, the events and milestones that we’ve been privileged to be a part of this year. Our 20th anniversary year is winding down and we are wrapping it up by spreading good news & gratitude throughout the season. Your financial donations and selfless volunteerism are what enable us to continue providing vital services to our GIST community.



