By Dr. Anette Duensing, University of Pittsburgh Cancer Institute, and Dr. Maria Debiec-Rychter, KU Leuven, Belgium, LRG Research Team
Although most GISTs initially respond well to treatment with imatinib mesylate, many patients unfortunately develop resistance to imatinib as well as other tyrosine kinase inhibitors, such as sunitinib and regorafenib over time. New therapeutic strategies are therefore urgently needed. In this article, we will describe how high-throughput drug-screening techniques can quickly identify new drugs for GIST patients by testing a large number of compounds at once. The focus lies on explaining some of the technical aspects of such analyses before presenting data from our recently published study. 1
Potential new drugs for GISTs need to be tested for efficacy in relevant cellular systems that at the same time allow for rapid analyses. Patient-derived GIST cell lines grown in vitro (Latin for “in glass”; i.e. in a tissue culture dish) are ideal for such studies. These cell lines are established from fresh tumor tissue shortly after surgery. A small piece of the tumor is minced very finely and then enzymatically dissociated into single cells (Figure 1). These are then covered with nutrient-rich liquid (“culture medium”) and incubated at 37°C (or 98.6°F, i.e. body temperature) in a regulated atmosphere with oxygen and carbon dioxide. This technique allows growth and expansion (division into more culture dishes) of cancer cells for a few weeks as so-called short-term cultures. Some of these cultures will develop into immortal cell lines that can be propagated indefinitely, even be frozen and brought back to life. The determinants of this process, however, are largely unknown. We therefore cannot predict, which cells will make it into a permanent line and which ones will not. Luckily, the LRG Research Team has either developed or is in possession of most GIST cell lines that are available worldwide. These include cell lines that are sensitive to treatment with imatinib as well as those that were derived from imatinib-resistant GISTs.
To test new drugs for their efficacy, GIST cells are treated with various compounds and are then monitored for their reaction to the drugs. Most importantly, it is determined whether the cells stop dividing or are even being killed. This can be done using various techniques that either involve biochemical assays, microscopic evaluation or automated measurements of intracellular enzymes. In comparison, normal, non-cancerous control cells are tested and should not be killed. Otherwise, a substance might be too toxic.
The above-mentioned experimental process is quite labor-intensive. To test a large number of drugs at the same time, cells can therefore be transferred to special culture dishes that are comprised of multiple “wells” (Figure 2). Each well serves as a mini test tube and is given a different drug. Standard well numbers of such culture plates are 96, 384 or 1536 (the latter two will need a robot for handling), meaning that nearly 100 drugs (or more) can easily be tested at once. Proliferation (rate of cell division) and apoptosis (cell death) are measured using colorimetric assays in an automated system (luminometer; Figure 2).
The results are evaluated using computational methods. They compare the test values to the “negative control” (no drug treatment) and a “positive control” (a drug that is known to kill the cells) that were included in the same experiment. A potentially effective compound has to be active beyond a certain threshold to be considered a “hit”. Results are also visualized by creating so-called heat maps (Figures 2,3), in which individual values are represented as colors and are sorted according to similarity. Using these methods, active compounds are identified and are then further evaluated with respect to their minimal effective dose and mechanism of action in GIST cell lines.
Before a drug that was identified in a drug screen is further investigated for potential treatment in humans, it needs to be tested in vivo (Latin for “in a living organism”). In the past decades, a number of murine GIST models have been developed and have become very valuable tools for such studies. Generally, these models can be divided into two categories: 1) genetically engineered mouse (GEM) models and 2) propagation of human tumor tissues in specific strains of mice (xenografts; i.e., a tissue graft from a different species; xeno, Greek for “foreign”).
In GIST GEM models (2,3), the genetic profile of the mice is artificially altered. These mice express a mutated Kit gene (equivalent to the KIT mutations seen in human GIST) in every cell of their body and develop GIST-like tumors in their bowels. However, due to their location in the abdominal cavity, therapeutic responses of the tumors are difficult to assess in vivo without the help of expensive imaging techniques. Moreover, these GEM models cannot fully reproduce the complexity of molecular abnormalities that exist in human tumors. They may also not accurately predict the therapeutic response of the human setting, because they are mouse tumors.
The discovery of mouse strains that do not reject human tissue (nude athymic mice and SCID mice) permits the efficient growth of established human cell lines or human tumor tissue explants directly obtained from patient biopsies underneath the skin of these animals (Figure 4). These reconstituted solid tumors will develop over the course of several weeks and can be excised, dissected into five mm cubes and transplanted into additional cohorts of mice once they have reached a certain size.
To test the response to therapeutic regimens in vivo, mice are grouped into so-called cohorts (12-16 animals), which are exposed to a given compound or combination of drugs (experimental groups) or not treated (control group) for a period of two to three weeks (Figure 4).4 The tumor size is measured regularly during this time (every 3 days), allowing tumor growth assessment under therapy. The well being of the animals and side effects of treatment are monitored continuously. In addition, it is determined whether tumor growth returns over a four to eight week period after treatment discontinuation (“tumor re-growth assay”). At the end of the experiment, mice are sacrificed and tumors are collected for microscopic (histopathologic) and biochemical evaluation.
For histopathology, tumor specimens are processed in the same way as surgical specimens in pathology (Figure 5, left panel). The tissue is preserved in fixative (formalin) and then embedded in wax blocks, from which very thin sections (4 μm) are cut and placed onto glass slides. These sections can then be stained by various methods to evaluate the response to therapy. The grade of histopathological response is evaluated through standard hematoxilin and eosin (H&E) staining and basic morphologic assessment of the tumor tissue. This includes the magnitude of tissue decay, degeneration, fibrosis and immune cell response. Measuring the effect of the treatment on the rate of proliferation and cell death may be facilitated by immunohistochemistry, a technique that uses specific antibodies to stain proteins of interest within a tissue section. Biochemical assays, such as immunoblotting, are used to detect specific proteins in samples of tumor tissue extracts (Figure 5, right panel).
A final step in the evaluation of a newly identified drug is to determine its mechanism of action. This is primarily being done through functional biochemical and microscopic assays in tissue culture models. But additional studies can also be carried out in the preserved tissue specimens from the xenograft experiments.
In our recent study (1), we tested a panel of 89 drugs that are already FDA-approved for cancer treatment. This collection of drugs is provided to researchers through the National Cancer Institute’s (NCI) Developmental Therapeutics Program. All 89 drugs were tested in triplicate in 5 GIST cell lines (2 imatinib-sensitive, 3 imatinib-resistant) and two leiomyosarcoma cell lines in comparison. Our screening experiments successfully identified two major drug classes as being very effective in GIST – even in imatinib-resistant cell lines (Figure 3). This was unexpected, because most of the drugs included in the collection were classical chemotherapeutic agents – drugs that are traditionally known as not being effective in GIST. However, this notion came from studies that had been conducted before a reliable diagnosis of GIST was possible (before 1999). Hence, a number of non-GIST malignancies may have inadvertently been included. Moreover, a systematic testing of chemotherapeutic agents in GIST had not been done, in part due to the rarity of the disease.
Drugs that our study identified as effectively killing GIST cells either belonged to so-called transcriptional inhibitors or inhibitors of a DNA repair enzyme called topoisomerase II. One drug of each class was chosen for further follow-up. The transcriptional inhibitor mithramycin A blocks the first step (gene transcription) in a cellular process that is needed to produce proteins, for example the mutant KIT molecules that drive the survival of GIST cells. We could show that mithramycin A kills GIST cells (at least in part) by depriving them of KIT. Inhibition of topoisomerase II by mitoxantrone leads to the introduction of so-called DNA double-strand breaks, the most dangerous type of DNA damage. Accumulation of too many of such breaks leads to cell death. Our experiments confirmed that this is also the case in GIST cell line models. In the next step, both drugs were tested in mouse xenografts where they showed significant anti-GIST efficacy.
In summary, our high-throughput screening study successfully identified two classes of FDA-approved cancer drugs with high efficacy in GIST cell line and in vivo models. Clinical trials are currently being discussed.
References
- Boichuk et al., Cancer Res 2014; 74:1200-1213.
- Sommer et al., Proc Natl Scad Sci USA 2003; 100:6706-6711.
- Rubin et al., Cancer Res 2005; 65:6631-6639.
- Floris et al., Clin Cancer Res 2009; 15:4066-4076.
Figures
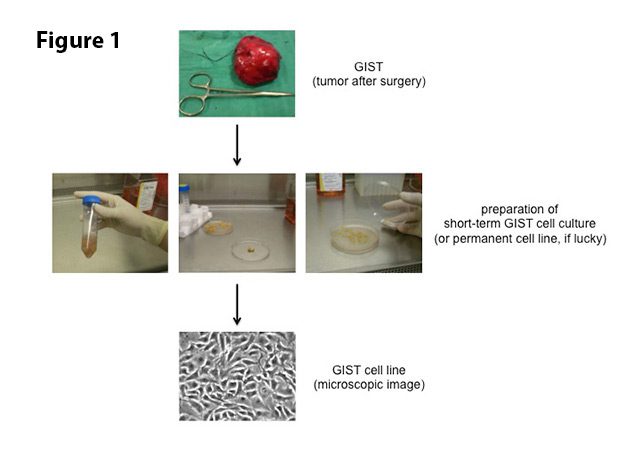
Figure 1. To establish a GIST cell line, pieces of a surgically removed GIST are transported to the laboratory in a sterile salt solution. Under sterile conditions, the tissue is minced into small pieces and enzymatically dissociated into single cells. Debris is removed by centrifugation and the cells are then covered with nutrient-rich liquid (“culture medium”). Incubation at body temperature (37°C/98.6°F) allows for growth and expansion of cells for a few weeks (short-term culture). Some of these cultures will develop into a permanent cell line.
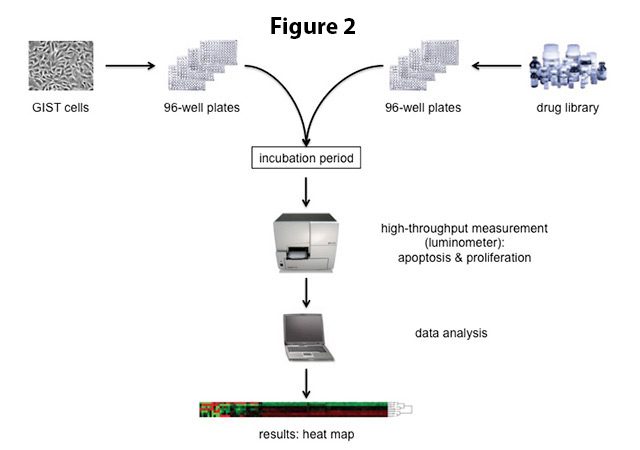
Figure 2. For high-throughput drug screening, GIST cells are transferred into 96-well plates. Each well serves as a mini-test tube that is treated with a different drug from a compound library. Cells are incubated with the respective drugs for a certain amount of time (usually 2-3 days) before the effect of the drugs on cell proliferation and apoptosis is measured in a luminometer. The data are analyzed and presented visually in a so-called heat map (bottom).
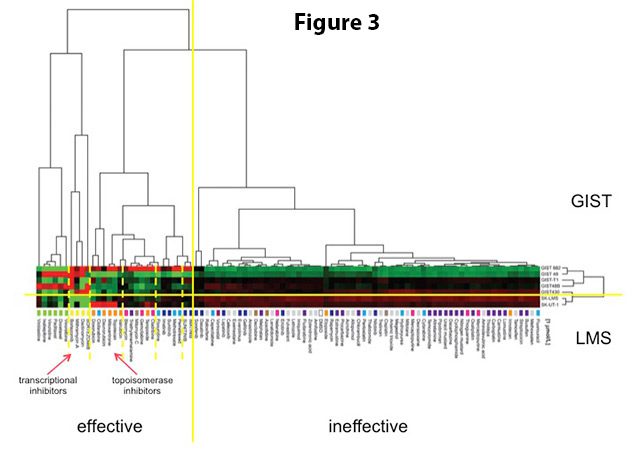
Figure 3. Efficacy of 89 FDA-approved cancer drugs in GIST vs. leiomyosarcoma (LMS) cells. 2 imatinib-sensitive and 3 imatinib-resistant GIST cell lines and 2 leiomyosarcoma cell lines were tested. Each row represents one cell line and each column represents one drug. Automated data analysis independently grouped results according to their similarity. Interestingly, all GIST cell lines and all LMS cell lines grouped together (divided by yellow horizontal line). Similarly, drugs that were effective in GIST versus those that were not also grouped together (divided by the yellow vertical line). Drugs also grouped according to their mechanism of action (divided by dashed yellow vertical lines). The most effective drug classes in GIST were transcriptional inhibitors and inhibitors of topoisomerase II (indicated by red arrows). Figure adapted from Boichuk et al, Cancer Research 2014.
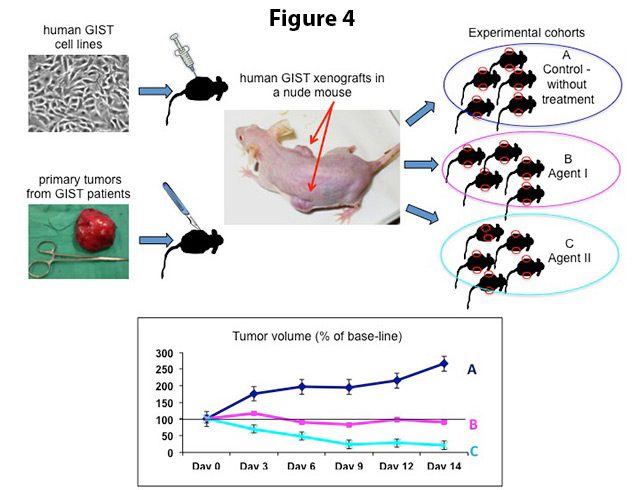
Figure 4. Upper part: Human GIST xenografts can be grown in nude mice either by injection of GIST cells from established cell lines or by bilateral insertion of fresh, surgically removed tumor fragments under the skin of the mouse. From the primary xenograft, tumors are re-transplanted into additional cohorts of mice for drug testing. Lower part: To assess the impact of treatment on tumor growth, the tumor volumes in non-treated (A, control) and experimentally treated cohorts (B and C) are monitored continuously until the end of the experiment.
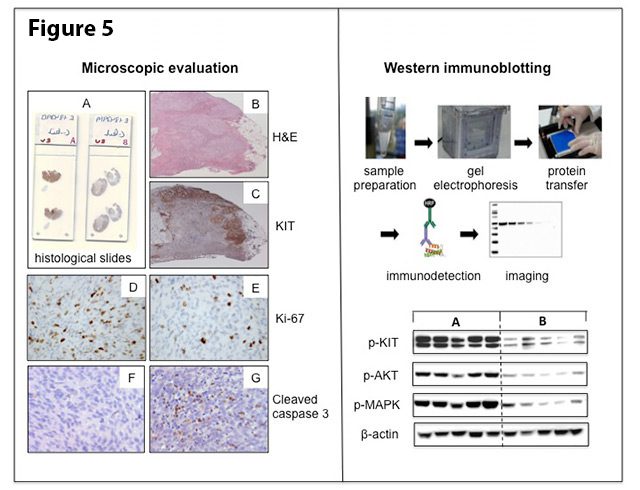
Figure 5. Histopathologic evaluation (left panel): Paraffin sections of tumor specimens on glass slides (A) are stained with H&E (B) and KIT immunostaining (C) for the morphologic assessment of the response to treatment. The KIT immunostaining (brown color) highlights the remaining tumor islands at the periphery of the tumor section. Marked reduction of the proliferative activity and the dramatic increase in apoptotic activity in treated (E and G) versus untreated (D and F) tumors, as evidenced by the number of Ki-67 and cleaved caspase 3 immunostained cells, respectively. Biochemical evaluation (right panel): Schematic presentation of the Western immunoblot technique (upper part) and an example of an immunoblot (lower part), showing differences in activation of KIT and its downstream signalling pathways in non-treated (cohort A) versus imatinib-treated tumors (cohort B). Each vertical lane reflects proteins isolated from one tumor sample.






